-
Viewpoint on 'IONS'
Viewpoint on 'Scientific Literacy'
- Proudly sponsored by
-

-
A Seriously Defocused Spider

A jumping spider about to pounce on its prey needs to be able to accurately measure its distance from its target in order to be able to hit it with precision. This is possible thanks to a visual approach previously unknown to exist in nature.
-
Glowing, Enchanted Materials

Never underestimate the magic of a glowing object. Luminescent materials have fascinated mankind for centuries. This enchantment continues with novel materials that can go on glowing for days without receiving continuous power from an energy source.
-
Rewinding Plasmons Back in Time
Day-to-day life common sense often does not apply in science. But sometimes it works better than any other approach. Scouts know that retracing back clear markers on the way can avoid getting lost, the same principle has been recently proved to work in nanoplasmonics.
Volume 11 Story 2 - 20/10/2010
If you had to bash a nail into the wall, you could probably use a stone to fulfill the task. Would you not agree, however, that a hammer would be a far more efficient tool to use? Similarly, when carrying out an experiment, researchers often look into a phenomenon with techniques that are good enough
but only just about. In fluorescent imaging, for example, it is easy nowadays to achieve a spatial resolution of a few nanometers. Now, the work carried out by Alexandros Pertsinidis and colleagues at the University of California Berkeley (California, USA) shows that fluorescent imaging can be improved to obtain subnanometer resolution.
According to a well-known paradigm in physics, objects separated by distances shorter than the limit of diffraction of visible light — approximately 250nm — cannot be distinguished with an ordinary optical microscope. In the last decade, different super-resolution imaging techniques based on fluorescence have been successful in overcoming this fundamental limit and have managed to achieve a resolution of a few nanometers [1]. Fluorescence is visible light spontaneously emitted by some molecules upon absorbing light of a shorter wavelength.
Wait a minute, though! If fluorescence is light, and if light is limited by diffraction, how can the resolution be better than 250 nanometers? All we need to do is use additional information about the fluorescence process in order to improve this resolution. On the one hand, two molecules which are separated by less than the limit of diffraction, and which emit light of the same color (wavelength) are obviously not distinguishable from one another. On the other hand, if they emit light in different instants of time, or of two different colors, we can locate the peak of emission for each of the molecules independently and measure distances of a few nanometers. "Although fluorescence based techniques feature significantly better performance — with a resolution of a few nanometers — than ordinary optical microscopy," explains Pertsinidis, "assemblies of biological molecules still proved too small to characterize." Odd as it may sound, a resolution of a few nanometers may not be enough when trying to shed light on processes happening on a molecular scale.
Can fluorescence microscopy be pushed to the next level of perfection? To improve the resolution of fluorescence microscopy even further, and increase it to less than a nanometer, a more careful characterization of its systematic sources of errors is needed. In a nutshell, as Pertsinidis points out, this is what the new proposal is about: "A sophisticated feedback system can carefully minimize systematic errors that are the result of the intrinsic imperfection of a microscope, its imaging optics and its CCD detector, and thus achieve another order-of-magnitude improvement. With our proposal, the separation between individual fluorescent molecules can be measured with a precision and accuracy of less than one nanometer, allowing us for the first time to characterize optically the arrangement of individual protein subunits within larger biomolecular complexes."
As a first attempt, this work focused on measuring the separation between two different molecules with fluorescence of different colors. "Previous approaches that used color as a separator to achieve sub-diffraction resolution had registration errors that ranged from between five to ten nanometers," Pertsinidis clarifies. "Our approach reduced this error to half a nanometer." First, it was essential to correct possible errors in measurement resulting from a non-uniform response of the different pixels in the CCD detector. A second challenge came about when we had to reduce the intrinsic internal motion of the imaged structures during each snapshot with the CCD (about one millisecond), thus reducing the localization uncertainty.
Pertsinidis and colleagues demonstrated the feasibility of this approach in two simple cases: firstly by manipulating a DNA molecule with an optical trap to fix its position and orientation, and then by tethering some trans-membrane proteins on a bio-compatible surface with multiple linkages. The fact that it is necessary to anchor the position of the molecules in order to take a snapshot of them with subnanometer resolution could represent a drawback when dealing with "molecules with a substantial inherent flexibility (such as disordered protein domains) and/or with large internal motions," as Pertsinidis explains. "However, there are many interesting cases where sub-diffraction optical imaging can be successfully applied."
The researchers are already working on ways of extending their technique to a more general configuration that does not rely on different colors for imaging subnanometer resolution. "This new approach can be used to perform measurements between single-molecule fluorescent probes with subnanometer precision and accuracy, while operating at room temperature and in physiological buffer conditions," Pertsinidis says. "This result is an important proof of principle that shows how our technique could be applied to further subnanometer measurements on more complex molecular arrangements. The ability to characterize biological structures with non-invasive optical means, and to resolve structures at the molecular level, will help expand our knowledge base in many important areas of molecular/cell biology and neuroscience."
For the researchers working in the field of single molecule detection, or in that of super-resolution fluorescent imaging, these results represent a beautiful refinement, to quote Harald Hess at the Howard Hughes Medical Institute (Maryland, USA). "It is nice to see a very careful, comprehensive and complete method described," he adds. "I believe these results fall into that category where attention to detail can really help pull the ultimate information out of a single molecule, thus offering a tool which will enable us to better understand and visualize biomolecular dynamics, such as the small conformational changes which proteins undergo when folding. Ultimately, how to make use of such a tool is open to the creativity of biologists and researchers."
[1] Niek van Hulst, Many Photons Get More out of Diffraction, Opt. Photon. Focus (2009) 4, 1. http://www.opfocus.org/index.php?topic=story&v=4&s=1

Subnanometer Perfection
According to French philosopher Voltaire (1694-1778) the perfect is the enemy of the good, thus not exactly desirable. As recent results show, however, where scientific research is concerned this may not always hold true.
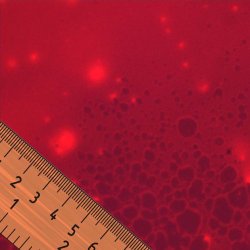
A subnanometer ruler for fluorescence. Different super-resolution imaging techniques based on fluorescence have been successful in measuring distances between fluorescent molecules that are only a few nanometers apart. By carefully characterizing systematic errors, it is possible to increase the resolution of these techniques to less than a nanometer.
According to a well-known paradigm in physics, objects separated by distances shorter than the limit of diffraction of visible light — approximately 250nm — cannot be distinguished with an ordinary optical microscope. In the last decade, different super-resolution imaging techniques based on fluorescence have been successful in overcoming this fundamental limit and have managed to achieve a resolution of a few nanometers [1]. Fluorescence is visible light spontaneously emitted by some molecules upon absorbing light of a shorter wavelength.
Wait a minute, though! If fluorescence is light, and if light is limited by diffraction, how can the resolution be better than 250 nanometers? All we need to do is use additional information about the fluorescence process in order to improve this resolution. On the one hand, two molecules which are separated by less than the limit of diffraction, and which emit light of the same color (wavelength) are obviously not distinguishable from one another. On the other hand, if they emit light in different instants of time, or of two different colors, we can locate the peak of emission for each of the molecules independently and measure distances of a few nanometers. "Although fluorescence based techniques feature significantly better performance — with a resolution of a few nanometers — than ordinary optical microscopy," explains Pertsinidis, "assemblies of biological molecules still proved too small to characterize." Odd as it may sound, a resolution of a few nanometers may not be enough when trying to shed light on processes happening on a molecular scale.
Can fluorescence microscopy be pushed to the next level of perfection? To improve the resolution of fluorescence microscopy even further, and increase it to less than a nanometer, a more careful characterization of its systematic sources of errors is needed. In a nutshell, as Pertsinidis points out, this is what the new proposal is about: "A sophisticated feedback system can carefully minimize systematic errors that are the result of the intrinsic imperfection of a microscope, its imaging optics and its CCD detector, and thus achieve another order-of-magnitude improvement. With our proposal, the separation between individual fluorescent molecules can be measured with a precision and accuracy of less than one nanometer, allowing us for the first time to characterize optically the arrangement of individual protein subunits within larger biomolecular complexes."
As a first attempt, this work focused on measuring the separation between two different molecules with fluorescence of different colors. "Previous approaches that used color as a separator to achieve sub-diffraction resolution had registration errors that ranged from between five to ten nanometers," Pertsinidis clarifies. "Our approach reduced this error to half a nanometer." First, it was essential to correct possible errors in measurement resulting from a non-uniform response of the different pixels in the CCD detector. A second challenge came about when we had to reduce the intrinsic internal motion of the imaged structures during each snapshot with the CCD (about one millisecond), thus reducing the localization uncertainty.
Pertsinidis and colleagues demonstrated the feasibility of this approach in two simple cases: firstly by manipulating a DNA molecule with an optical trap to fix its position and orientation, and then by tethering some trans-membrane proteins on a bio-compatible surface with multiple linkages. The fact that it is necessary to anchor the position of the molecules in order to take a snapshot of them with subnanometer resolution could represent a drawback when dealing with "molecules with a substantial inherent flexibility (such as disordered protein domains) and/or with large internal motions," as Pertsinidis explains. "However, there are many interesting cases where sub-diffraction optical imaging can be successfully applied."
The researchers are already working on ways of extending their technique to a more general configuration that does not rely on different colors for imaging subnanometer resolution. "This new approach can be used to perform measurements between single-molecule fluorescent probes with subnanometer precision and accuracy, while operating at room temperature and in physiological buffer conditions," Pertsinidis says. "This result is an important proof of principle that shows how our technique could be applied to further subnanometer measurements on more complex molecular arrangements. The ability to characterize biological structures with non-invasive optical means, and to resolve structures at the molecular level, will help expand our knowledge base in many important areas of molecular/cell biology and neuroscience."
For the researchers working in the field of single molecule detection, or in that of super-resolution fluorescent imaging, these results represent a beautiful refinement, to quote Harald Hess at the Howard Hughes Medical Institute (Maryland, USA). "It is nice to see a very careful, comprehensive and complete method described," he adds. "I believe these results fall into that category where attention to detail can really help pull the ultimate information out of a single molecule, thus offering a tool which will enable us to better understand and visualize biomolecular dynamics, such as the small conformational changes which proteins undergo when folding. Ultimately, how to make use of such a tool is open to the creativity of biologists and researchers."
[1] Niek van Hulst, Many Photons Get More out of Diffraction, Opt. Photon. Focus (2009) 4, 1. http://www.opfocus.org/index.php?topic=story&v=4&s=1
Giorgio Volpe
2010 © Optics & Photonics Focus
GV is currently working on his doctoral thesis at ICFO - The Institute of Photonic Sciences, Barcelona (Spain).

Alexandros Pertsinidis, Yunxiang Zhang & Steven Chu, Subnanometre single-molecule localization, registration and distance measurements, Nature (2010) 466, 647-651 (link).