-
Viewpoint on 'IONS'
Viewpoint on 'Scientific Literacy'
- Proudly sponsored by
-

-
EPS QEOD Thesis Prizes: The Winners of 2011

Six young scientists have received the prestigious QEOD Thesis Prize, awarded by the Quantum Electronics and Optics division (QEOD) of the European Physical Society (EPS). Congratulations!
-
A Seriously Defocused Spider

A jumping spider about to pounce on its prey needs to be able to accurately measure its distance from its target in order to be able to hit it with precision. This is possible thanks to a visual approach previously unknown to exist in nature.
-
Plasmons: Transform!
Latest experiments demonstrate an easy way to freely transform electromagnetic waves moving on metals. This yields promise of powerful applications, ranging from computers and biomedical devices, to solar cells and cell phones.
Decoding the Code of Life
Reading information stored by genes — also known as gene sequencing — is a vital task to the study of life itself. A radically new technology has set the stage for a revolution in the deciphering of DNA strands.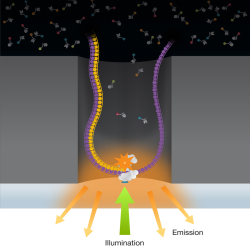
A zero mode waveguide used for real time sequencing of a DNA molecule. A single molecule of DNA bound to DNA polymerase is immobilized at the bottom of a zero mode waveguide, which is illuminated from below by laser light. The color of the emitted fluorescence indicates the present nucleotide being added to the chain.
Sometimes delicacy of a flower is life,
Sometimes hardness of life is life,
Desire to get something is life,
Sadness to loose something is life,
Sometimes a sweet dream is life,
Sometimes it looks punishment is life,
I don`t uptill now,
What is life?
— Dr. Ram Sharma
What is life? — Be it scientific or philosophical, there is no simple answer to this question. When taking the scientific approach, hints can come from the information stored in the code of life, the genes. The process of decoding the information in the genes is termed gene sequencing, or DNA sequencing. This is the reason why the human genome project, which aimed to sequence about 25,000 human genes, has been heralded as a major milestone in science. However, the currently existing techniques for gene sequencing are too slow and also suffer form various other drawbacks. Now a radically new approach proposed by the California-based company Pacific Biosciences promises to speed up this process to unprecedented levels.
Drawing an analogy to modern day computers, a cell, the functional unit of life, has hardware the cellular machinery and software the genetic code. Such software describes both the gears of the cellular machinery and the interactions among such gears as the living system persists through time [1]. Just as the bits store digital information in a computer in the form of 1s and 0s, the DNA codes information in its nucleotide bases: adenine (A), guanine (G), cytosine (C), and thymine (T). Continuing with the analogy, as a set of 8 bits forms a byte, a set of 3 nucleotides forms a codon that codes for a specific amino-acid. Finally, a set of codons forms a gene and codes a protein, which is a long chain of amino-acids. A chromosome, which is a single piece of coiled DNA, contains a large number of genes and an organism has a number of chromosomes, coding for a wide variety of proteins that give each cell its meaning and identity. Human beings, for example, have 23 pairs of chromosomes, with about 25,000 genes, and over 6 billon nucleotides.
DNA sequencing decodes the order in which the nucleotide bases (A, T, G and C) appear in a fragment of DNA. One of the most widely used techniques for gene sequencing is the Sanger method, which exploits the way in which DNA replicates. "The Sanger sequencing method is limited to a few tens of thousands of bases of sequences per hour, and the cost to sequence the human genome is millions of dollars," notes Stephen Turner of Pacific Biosciences. "A second generation of sequencing technologies has arrived that dramatically increases the throughput the number samples that can be sequenced in a given time and reduces the cost of sequencing. However, these techniques have reduced readlength and take longer to complete than the Sanger technique."
A DNA molecule consists of two long chains of nucleotides. The two chains are complementary: to each A in the first chain corresponds a T in the second chain, and to each C a G, and vice versa. Such structure allows for replication by separating the two chains and adding the complementary nucleotides to each of the two unwound chains. In the Sanger method, 4 DNA replication reactions occur in parallel. To each reaction is then added only one of four dideoxynucleotides, which are modified nucleotides with the property that the DNA replication process stops as soon as one of them gets added instead of a nucleotide. Since each reaction involves replication of a large number of DNA molecules, the random chain terminations result in DNA fragments of varying lengths. A gel electrophoresis process for each of the four reactions then separates out the newly synthesized fragments by length, thus indicating the position of each nucleotide in the DNA fragment. This technique, however, does not provide real time read out of the sequencing data during DNA replication and is very time consuming and expensive.
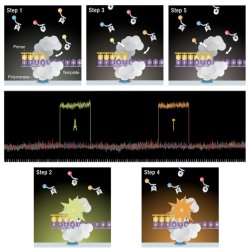
The process of real time DNA sequencing. (Step 1) A nucleotide forms an association with the template in the polymerase active site, (Step 2) causing an elevation of the fluorescence output on the corresponding color channel. (3) The fluorescent tag is liberated and diffuses out of the ZMW, thus ending the fluorescence pulse. (4) The polymerase translocates to the next position, and (5) the next cognate nucleotide binds the active site beginning the subsequent pulse.
This technology is a significant breakthrough in gene sequencing. "While the Sanger sequencing could read around 800 consecutive bases in around an hour, our sequencing method exceeds Sangers in readlength and can complete sequencing runs in under 30 minutes, all the while retaining the high-throughput associated with second generation systems," clarifies Turner. "This work describes a fundamental departure from all previously used DNA sequencing approaches," observes Harold G. Craighead from the School of Applied and Engineering Physics at Cornell University and cofounder of Pacific Biosciences. "Single molecule observation allows for the speed in the sequencing of a DNA molecule to progress at the rate of the polymerase enzyme activity, unlimited by any macroscopic fluid handling or chemistry. By engineering many polymerase molecules into an ordered array, they act as independent sequencers operating in parallel. This points to a dramatic increase in rapidly-available low-cost sequence data and its broader use in research and medical applications."
There have been many breakthroughs achieved in this work. An especially important one is the use of nanophotonic structures the zeromode waveguide (ZMW) to observe a single DNA molecule. A high concentration of nucleotides is required by the DNA polymerases that catalyze the process of DNA replication. Since the goal was to observe a single molecule of DNA such high concentrations could only be maintained by significant reduction in the observation volume, that was far beyond what was offered by conventional techniques like confocal microscopy. "We invented a nanophotonic observation device called the zeromode waveguide to overcome this problem," explains Turner. "Zeromode waveguides are nanometer-scale holes in a metal film. Laser illumination energy that impinges on the hole is blocked from passing through because the hole acts like a waveguide in cutoff." The ZMW enables an observation volume in the zeptoliter (10-21 liter) regime, allowing for the first time observation of single-molecules at concentrations relevant to polymerases.
A special surface chemistry is used to immobilize a single molecule of DNA template-bound DNA polymerase at the bottom of the ZMW. The new technique uses specially engineered nucleotides that carry a fluorescent tag. These fluorescently tagged nucleotides are then used as the building blocks in the DNA replication process.
As the replication progresses, nucleotides are added one after the other to the newly formed strand. There is a separate tag for each of the four nucleotides. As one nucleotide is added to the chain, it gives a fluorescent signal indicative of its type in the detection volume. Hence, by looking at the color of the fluorescence, it can be determined what type of nucleotide has been added. This is like each nucleotide calling out its identity as it gets added to the chain. Therefore, by looking at the color of the fluorescence, it is determined what type of nucleotide has been added.
Due to a very novel design, the fluorescent tag is released and gets diffused away from the detection volume as soon as the bonding process of the nucleotide to the chain is completed. "This means that fluorescent labels are visible during catalysis, but are spontaneously clipped away afterwards, leaving behind completely natural DNA synthesis product," points out Turner. "This is in stark contrast with base-labeling techniques where the fluorescent label remains behind hindering polymerization and causing accumulation of fluorescence background." The end result is a fluorescent pulse that lasts only as long as it takes to complete the bonding process. As the next nucleotide gets added, the detection volume is free form the influence of the previous nucleotide fluorescence tag. The whole sequencing process is carried out in real time.
The system enables simultaneous detection of fluorescence from 3000 ZMWs, which means that 3000 independent sequencings can be preformed in parallel. Since there are four fluorescent tags the excitation is provided by a multilaser line consisting of two laser wavelengths. These two wavelengths are sufficient enough to excite the four possible fluorescent tags. The simultaneous excitation of 3000 ZMWs is enabled by holographically splitting a multilaser line into 3000 beamlets. "A polymerase molecule immobilized in the bottom of the hole is illuminated by an evanescent field that extends a few nanometers into the ZMW before dying out," states Turner. Since the detection aslo needs to be done simultaneously from 3000 ZMWs, a confocal detection system with a 3000 multiplexed pinhole array is employed. A prism based system is used to discriminate between the four fluorescent colors. "Through the use of a holographic phase mask and a confocal pinhole array we create thousands of confocal imaging volumes that give sensitive detection at the same time as rejecting out-of-focus noise in the system," remarks Turner.
One of the standing goals of biomedical research is to fully decode the information in genes and to understand how mutations in these genes lead to diseases. "Because of the complexity of the genome it will require many thousands if not millions of individuals to be fully sequenced before we fully understand the relationship between genes and diseases," says Turner. "This technology will speed up the arrival of that understanding by reducing the time and costs associated with sequencing."
This technology is truly groundbreaking. To give you an idea of exactly how remarkable this gene sequencing technology really is to geneticists, you can compare it to upgrading from a bicycle to a sport car. It may not be too far-off in the future when scientists, armed with technologies such as these, are finally able to decipher every bit of the genetic code in order to answer the question: What is life?
[1] Daniel E. Koshland Jr., The Seven Pillars of Life, Science 295, 2215-2216 (2002).
Manoj Mathew
2009 © Optics & Photonics Focus
MM is currently working on his doctoral thesis at ICFO - The Institute of Photonic Sciences, Barcelona (Spain).

John Eid et al., Real-Time DNA Sequencing from Single Polymerase Molecules, Science (2009) 323, 133-138 (link).